Scalable Research Achievement
SSR marker technology package for variety identification of white Guinea Yam
Description
White Guinea yam (Dioscorea rotundata), one of the most important cash crop for farmers and a major staple food for the people of West Africa, retains huge potential for alleviating widespread poverty and hunger in the region. Now, yam research is at a turning point, and recent advancements in genetics and mass propagation are expected to boost breeding efficiency and dissemination of improved varieties. On the other hand, since it is difficult to distinguish varieties based on the visible characteristics of the shoot and tuber of yam (Fig. 1), mechanical mixture between varieties grown in the same field has been a serious problem through all the steps in the breeding and propagation process, including planting, cultivation, harvesting, and storage. To overcome this problem, a simple tool to identify varieties was highly desired.
To enable variety identification of white Guinea yam, a Simple Sequence Repeat (SSR) marker system was adopted based on its various advantages such as high reproducibility, low cost, and high polymorphism. In the initial setup, 16 SSR markers that can be used to distinguish varieties and genetic resources effectively were selected from the 90 SSR markers developed in our previous study (JIRCAS Research Highlights 2015, B05) (Fig. 2). Additionally, various tools to enable successful variety identification using the developed SSR markers and maximize the benefits to users in the breeding and seed sectors were subsequently developed. For example, the developed web application “Minimum SSR Marker Finder for Guinea yam” (https://www.jircas.go.jp/en/database/yam_toolkit/finder) linked with the database contains SSR polymorphism data of over 550 varieties and lines (as of February 2020), and supports identification of the minimum set of SSR markers that can distinguish the varieties selected by each user. In addition to the conventional DNA extraction method using a young leaf sample, we developed the “Sample collection and DNA extraction methods for tuber skin” to widen the user’s choice of period to conduct variety identification (Fig. 3). Also, the proposed “Sample bulking method” specially designed for large-scale propagation of yam seed tubers, enables the yam seed sector to reduce time and cost in quality control and quality assurance (Fig. 4).
These useful tools and methods were assembled as an “SSR marker technology package for variety identification of white Guinea yam” to cover all necessary steps to distinguish varieties (Fig. 5), and the website titled “Yam variety identification toolkit” (https://www.jircas.go.jp/en/database/yam_toolkit) was launched to support users in conducting variety identification with various guidance, tips, manuals, videos, and images. The technology package can reduce time and cost, and provide flexibility for users to ensure the uniformity of the materials before planting, prevent mechanical mixture in various stages of cultivation, and assure the quality of the products. We expect this technology package to support seed growers, extension officers and inspection officers, as well as yam researchers involved in various stages of yam improvement and dissemination. This in turn could help boost breeding efficiency and dissemination of improved varieties, and further improve food security and livelihood in West Africa.
Figure, table
-
Fig. 1. White Guinea yam (D.rotundata)
Left: Shoots of multiple varieties grown in a farmer’s field
Right: Tubers obtained from a single plant (DrDRS-139) -
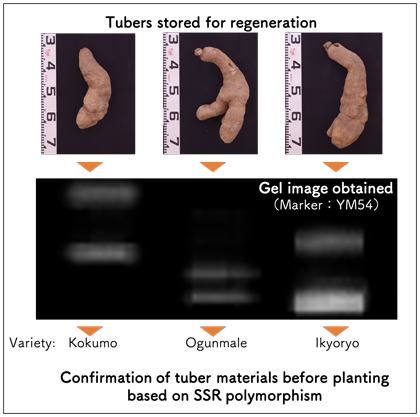
Fig. 2. Identification of varieties with similar tuber shapes using SSR markers
-
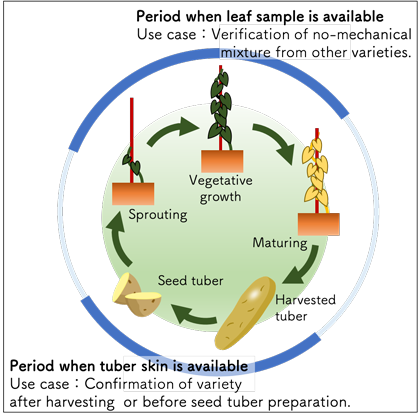
Fig. 3. Expanded period for variety identification with two types of samples
-
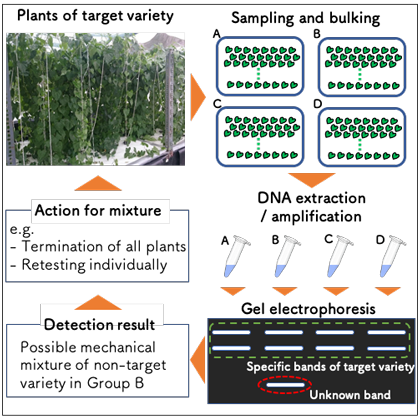
Fig. 4. Utilization of the sample bulking method to maintain purity of target variety
-
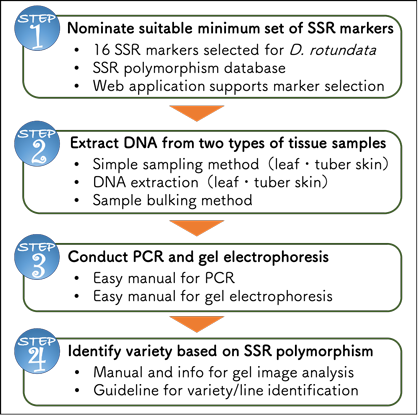
Fig. 5. SSR marker technology package for variety identification of white Guinea yam
- Affiliation
-
Japan International Research Center for Agricultural Sciences Crop, Livestock and Environment Division
Japan International Research Center for Agricultural Sciences Tropical Agriculture Research Front
- Classification
-
技術
- Research project
- Program name
- Term of research
-
FY2019 (FY2011-FY2020)
- Responsible researcher
-
Muranaka Satoru ( Crop, Livestock and Environment Division )
Yamanaka Shinsuke ( Tropical Agriculture Research Front )
MIERUKA ID: 001785Tamiru Muluneh ( Iwate Biotechnology Research Center )
Agre Paterne ( International Institute of Tropical Agriculture )
- ほか
- Publication, etc.
-
https://doi.org/10.2135/cropsci2014.10.0725
Tamiru M et al (2015) Crop Science, 55:2191–2200
Yam variety identification toolkit https://www.jircas.go.jp/en/database/yam_toolkit
- Japanese PDF
-
2019_B03_A4_ja.pdf805.61 KB
2019_B03_A3_ja.pdf323.03 KB
- English PDF
-
2019_B03_A4_en.pdf590.26 KB
2019_B03_A3_en.pdf1.09 MB
- Poster PDF
-
2019_B03_poster_fin.pdf336.54 KB