Characterization of Erianthus arundinaceus collected from Japan based on nuclear DNA content and simple sequence repeat markers
Description
Erianthus arundinaceus, a member of the Saccharum complex, has been receiving increased attention as a bioenergy crop and a potential resource for sugarcane improvement. Although Erianthus species are widely distributed in tropical and subtropical climates, some wild accessions are well adapted to temperate Japan. To our knowledge, these are wild E. arundinaceus accessions that have adapted to the northernmost regions, and thus are potentially novel genetic resources for the development of cultivars or breeding lines suitable for growth in the temperate zone. However, little is known about the genetic characteristics of the Japanese accessions. To facilitate genetic studies and breeding programs, we developed simple sequence repeat markers from E. arundinaceus and obtained basic knowledge about the genetic background of this species collected in Japan.
We investigated the 2C DNA content and SSR polymorphisms in E. arundinaceus accessions collected from three climatic zones (temperate, subtropical, and tropical) in Japan and Indonesia with flow cytometric analysis. All examined accessions fell into two groups reflecting collection location: the temperate zone in Japan (mean 8.06 pg) and the subtropical and tropical zones in Japan and Indonesia (mean 7.56 pg). Although DNA content differed significantly between the accessions from temperate and tropical/subtropical zones, chromosome number was inferred to be identical in all accessions. A phylogenetic tree of 29 accessions based on detected fragments from 39 SSR primer pairs classified Japanese (26 accessions) and Indonesian (3 accessions) accessions into well-defined distinct groups. Both 2C DNA content and phylogenetic analysis subdivided Japanese accessions into two groups, suggesting that some of the Japanese accessions had different genetic characteristics from other accessions from Japan and Indonesia. In addition, of the 39 SSR primer pairs, 31 primer pairs were amplified in other genera of the Saccharum complex including Saccharum spp., Miscanthus sinensis and Narenga porphyrocoma (Fig. 3).
It is unclear how the hexaploidy of Erianthus was established in evolution, but the significant differences in DNA contents between the two E. arundinaceus groups might reflect a geographical distribution and ecological adaptation after ancestral hexaploidization. Further comprehensive studies including additional genetic resources from other countries and genomic information are needed to fully understand the basis of DNA content variability in E. arundinaceus.
Figure, table
-
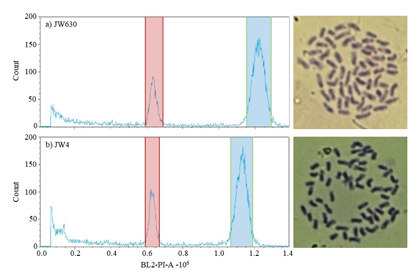
Fig. 1. Histograms of relative DNA content and mitotic metaphase chromosome preparations of Erianthus arundinaceus from temperate (a) and subtropical (b) zones.
The peaks marked in red and blue were detected in the internal control and the experimental samples, respectively. The x-axis shows the number of propidium iodide (PI)-labeled cells detected using a BL2A filter (488 nm). The y-axes are on cell counts. -
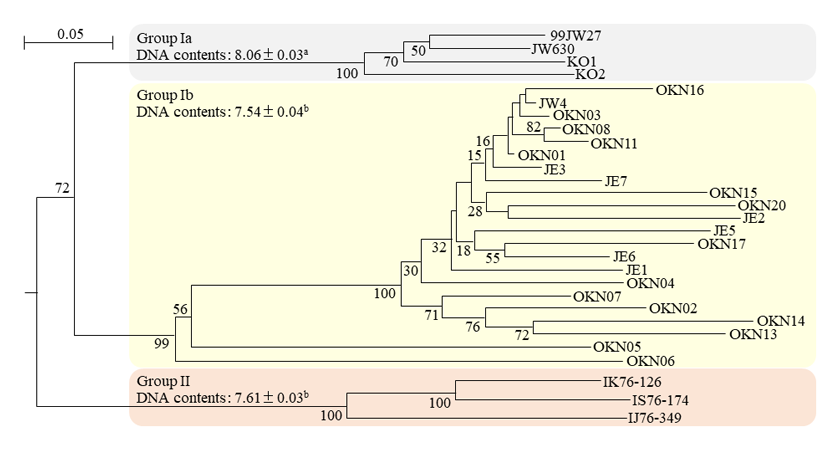
Fig. 2. Phenogram of the 29 accessions based on SSR genotyping data from 39 primer pairs.
Numbers beside the branches indicate the values (%) of 1,000 bootstrap replicates. Values of DNA content are mean ± standard deviation within each group. DNA content with different letters indicate statistically significant differences among groups (p<0.001). -
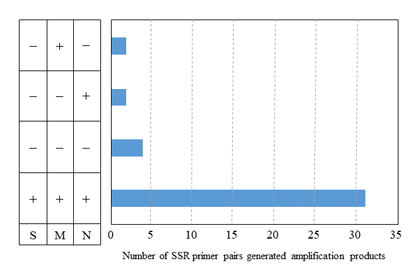
Fig. 3. Application of 39 SSR primer pairs to the Saccharum complex.
S: Saccharum spp. (7 species), M: Miscanthus sinensis (5 accessions), N: Narenga porphyrocoma (1 accession). Successful amplification (+), No amplification (–).
- Affiliation
-
Japan International Research Center for Agricultural Sciences Tropical Agriculture Research Front
- Research project
- Program name
- Term of research
-
FY2017(2011~2020)
- Responsible researcher
-
Tsuruta Shin-ichi ( Tropical Agriculture Research Front )
Teraiima Yoshifumi ( Tropical Agriculture Research Front )
Ebina Masumi ( Institute of Livestock and Grassland Science, NARO )
Kobayashi Makoto ( Institute of Livestock and Grassland Science, NARO )
MIERUKA ID: 000933Takahashi Wataru ( Institute of Livestock and Grassland Science, NARO )
- ほか
- Publication, etc.
-
https://doi.org/10.1007/s11032-017-0675-z
Tsuruta S et al. (2017) Mol Breed, 37
https://doi.org/10.1007/s11738-017-2519-1Tsuruta S et al. (2017) Acta Physiol Plant, 39
- Japanese PDF
-
A4 312.87 KB
A3 336.81 KB
- English PDF
-
A4 161.37 KB
A3 150.14 KB
- Poster PDF
-
2017_B11_poster.pdf465.86 KB