Identification of cis-acting promoter elements in cold- and dehydration-induced transcriptional pathways in rice, soybean and Arabidopsis
Description
Low temperature and dehydration affect plant growth and productivity. Many genes respond to both stressors at the transcriptional level, and their gene products function in terms of stress tolerance and response. These genes include key metabolic enzymes, late embryogenesis-abundant (LEA) proteins, detoxification enzymes, chaperones, protein kinases, and transcription factors. The cis-acting elements that function in stress-responsive gene expression have been analyzed to elucidate the molecular mechanisms of gene expression in response to these stresses. The dehydration-responsive element (DRE) containing the core sequence A/GCCGAC is a cis-acting element that regulates cold- and dehydration-responsive gene expression in Arabidopsis (Arabidopsis thaliana). The abscisic acid (ABA)-responsive element (ABRE) containing the core sequence ACGTGG/T is a cis-acting element that regulates dehydration- and high salinity-responsive gene expression in Arabidopsis and rice (Oryza sativa). ABA-responsive gene expression requires multiple ABREs or an ABRE with a coupling element as a functional promoter.
In this study, oligo microarrays were used to identify cold- and dehydration-responsive genes in rice, soybean and Arabidopsis. The observed frequencies of all (46 = 4096) hexamer sequences in cold- and dehydration-inducible promoters were compared with standardized promoters to estimate conserved sequences and to determine representative cold- and dehydration-responsive transcriptional pathways in rice, soybean and Arabidopsis.
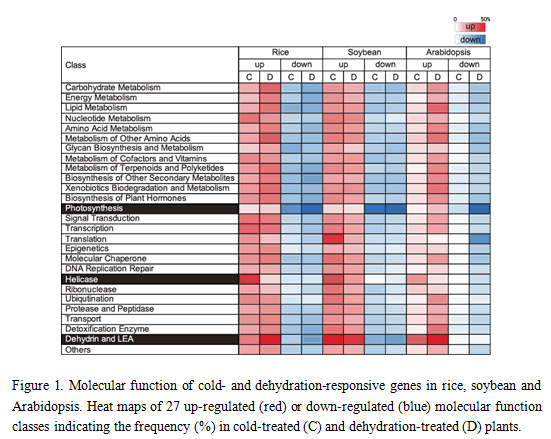
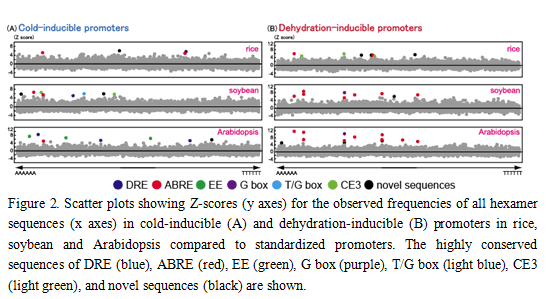
Microarray analyses were performed using the three species (rice, soybean and Arabidopsis) and the characteristics of identified cold- and dehydration-responsible genes were compared. Transcription profiles of the cold- and dehydration-responsive genes were similar among these three species, showing representative up-regulated (dehydrin/LEA) and down-regulated (photosynthesis-related) genes. All (46 = 4096) hexamer sequences in the promoters of the three species were investigated, revealing the frequency of conserved sequences in cold- and dehydration-inducible promoters. A core sequence of the ABRE was the most conserved in dehydration-inducible promoters of all three species, suggesting that transcriptional regulation for dehydration-inducible genes is similar among these three species with the ABRE-dependent transcriptional pathway. In contrast, the highly conserved sequences in cold-inducible promoters of Arabidopsis are different from those of rice and soybean. DRE is the most conserved sequence in cold-inducible promoters of Arabidopsis, but not in those of rice and soybean. The novel sequence and ABRE are the most conserved sequences in cold-inducible promoters of rice and soybean, respectively. In cold-inducible promoters, the conserved hexamer sequences were diversified among these three species, suggesting the existence of diverse transcriptional regulatory pathways for cold-inducible genes among the species.
Figure, table
- Affiliation
-
Japan International Research Center for Agricultural Sciences Biological Resources and Post-harvest Division
- Classification
-
Administration A
- Research project
- Program name
- Term of research
-
FY 2011 (FY 2011-FY 2015)
- Responsible researcher
-
Maruyama Kyounoshin ( Biological Resources and Post-harvest Division )
Shinozaki Kazuko ( University of Tokyo )
ORCID ID0000-0002-0249-8258 - ほか
- Publication, etc.
-
Maruyama et al. (2011) DNA Research (2012) 19: 37-49
- Japanese PDF
-
2011_07_A4_ja.pdf135.95 KB
- English PDF
-
2011_07_A4_en.pdf117.08 KB