Identification of a gene that promotes ammonium nitrogen (NH4+-N) uptake in rice
Description
Nitrogen is the most essential nutrient to plant growth and grain yield. There are two forms of inorganic nitrogen available to plants: one is NO3--form and the other is NH4+-form. NH4+-form is the major form in paddy fields, thus rice plants grown in the fields uptake mainly NH4+-form nitrogen as nitrogen source in roots. Therefore, promoting NH4+-form uptake is one of the major targets to increase grain yield in rice. It is known that NH4+ influx into roots by a high-affinity transport system (HAT) is down-modulated in response to elevated NH4+ concentration around soil surface. We hypothesized that canceling the down-modulation of NH4+ influx by HAT would be beneficial on the uptake of NH4+-form nitrogen in rice. However, no gene that modulates NH4+ influx by HAT has been identified in rice. In this study, we aimed to identify the gene that modulates HAT of rice roots with gene expression analyses. Furthermore, we tried to isolate a gain-of-function gene to promote uptake of NH4+-form nitrogen with rice Tos17 insertion mutants.
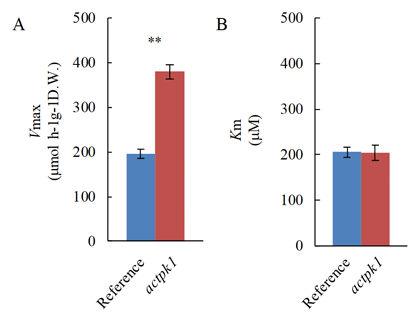
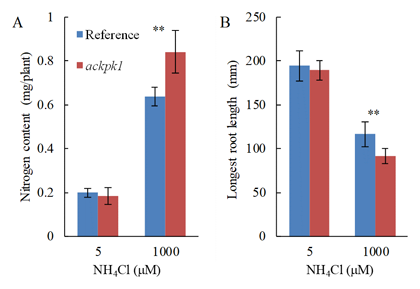
To isolate candidate genes concerned in down-modulation of HAT in roots of rice, we performed four-biological-repeat transcriptome analyses. A total of 28,381 out of 36,444 filtered probes were selected as differentially expressed genes based on a false discovery rate of ≤ 0.05. A strong candidate gene for down-modulation of HAT, the coding protein kinase gene OsACTPK1, showed 1,071 times higher expression in roots under NH4+-rich condition as compared with NH4+-deficient condition (Table 1). We then analyzed the detail of Tos17-inserted mutant of OsACTPK1 (actpk1 mutant) to elucidate OsACTPK1 as down-modulator of HAT. Kinetic analyses of NH4+ influx by HAT revealed that Vmax value of actpk1 mutant was 2 times higher than that of wild type, whereas there was no significant difference of Km value between these two lines (Fig. 1). These results indicated that down-modulation of HAT was canceled in actpk1 mutant due to the loss-of-function OsACTPK1. Total nitrogen content of the actpk1 mutant was significantly higher (+32%) than that of wild type in 1,000 mM NH4Cl condition (NH4+-rich condition), while the significant difference was not observed in 5 mM NH4Cl condition (NH4+-deficient condition) (Fig. 2A). Furthermore, the longest-root length of the actpk1 mutant was significantly lower (-22%) than that of wild type in 1,000 mM NH4Cl condition, while the significant difference was observed in 5 mM NH4Cl condition (Fig.2B).
We concluded that OsACTPK1 was a down-modulator of HAT and that loss-of-function ACTPK1 (actpk1 gene) could maintain activity of NH4+ influx even in elevated NH4+ concentration. The actpk1 gene could be effective in improving nitrogen use efficiency in a rice molecular breeding program. Also, reduction of root elongation in actpk1 mutant would be used as phenotypic marker in the program. Further analyses to characterize nitrogen use and grain yield of actpk1 are required for the program since OsACTPK1 would function to avoid NH4+ toxicity.
Figure, table
-
Rice Oligo DNA Microarray (4X44K RAP-DB) was used in this research. Total RNA was extracted from roots of rice plants grown for 10 days in 5 and 1,000 M NHTable 1. Details for the OsACTPK1 gene, a strong candidate for down-modulation of HAT Item Description Increase of gene expression in response to incease of external nitrogen concentration 1,071-fold RAP ID Os02g0120100 Protein function protein kinase -
Fig. 1. Kinetic properties of HAT in actpk1 mutant.Vmax value (A) and Km (B) to NH4+ were expressed in blue column for reference, Nipponbare and red column for actpk1. Plants were hydroponically grown for 10 days in 1,000 mM NH4Cl. Mean value with standard error (n=3-6) was plotted. Asterisks indicate probability of less than 0.01 between reference and actpk1 (ANOVA).
-
Fig. 2. Total nitrogen content (A) and longest root length (B)Plants were hydroponically grown for 10 days in two NH4+ conditions, 5 mM as deficient and 1,000 mM as rich condition of NH4+. Mean value with standard deviation (n=6 for total nitrogen content, n=14 for longest root length) was expressed in blue column for Nipponbare as reference and red column for actpk1, respectively. Asterisks indicate probability of less than 0.01 between reference and actpk1 (ANOVA).
- Affiliation
-
Japan International Research Center for Agricultural Sciences Biological Resources and Post-harvest Division
- Research project
- Program name
- Term of research
-
FY2017(2013~2020)
- Responsible researcher
-
Obara Mitsuhiro ( Biological Resources and Post-harvest Division )
Beier Macel Pascal ( Tohoku University )
Hayakawa Toshihiko ( Tohoku University )
KAKEN Researcher No.: 60261492 - ほか
- Publication, etc.
-
https://doi.org/10.1111/tpj.13824
Beier M et al. (2018) The Plant Journal, 93:992-1006
- Japanese PDF
-
A4 261.77 KB
A3 250.14 KB
- English PDF
-
A4 124.63 KB
A3 129.14 KB
- Poster PDF
-
2017_B05_poster.pdf331.83 KB