Variation at the Pup1 locus within the genus Oryza predates domestication
Description
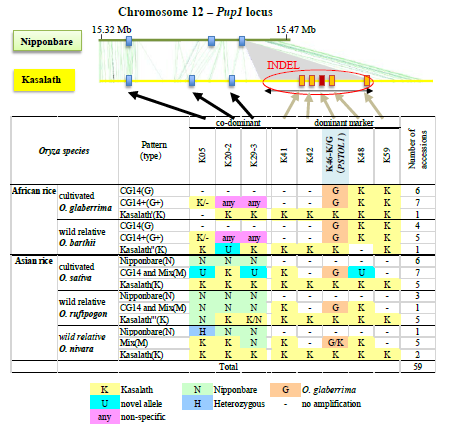
The deficiency of phosphorus (P) in soil is a worldwide problem, and though there are many approaches to tackle this problem, the development of rice cultivars with enhanced P efficiency would represent a sustainable strategy to improve the livelihood of resource-poor farmers. Recently, the Pup1 locus (Fig. 1), a major QTL for tolerance to P deficiency, was successfully narrowed down to a single-candidate gene, the protein kinase: P starvation tolerance (OsPSTOL1). The aim of this study was to search for novel OsPSTOL1 alleles and to survey Pup1 locus variation in Asian (O. sativa)- and African (O. glaberrima)-cultivated rice and their wild progenitors. This information would help in designing a suitable strategy for marker-assisted introgression of Pup1/PSTOL1 into rice megavarieties.
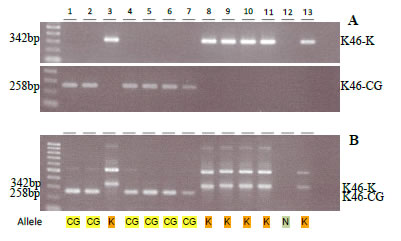
A novel OsPSTOL1 allele was detected in O. glaberrima. This allele has 35 base-pair changes (when aligned to Kasalath allele), but none of the functional domains were affected and it is expressed. Allele-specific markers were then developed for single PCR and/or duplex PCR system, which produce a band pattern clearly distinguishable on agarose gels (Fig. 2), and are therefore suitable for most marker laboratories throughout the world. Using these markers to survey allelic distribution of PSTOL1 across the genus Oryza showed that the novel allele is common in accessions belonging to O. glaberrima and its ancestor O. bartii, but is not restricted to African rice as O. sativa, O. rufipogon, and O. nivara accessions do carry the glaberrima allele at low frequency (Fig. 1).
Using additional allele-specific markers across the entire Pup1 locus revealed two main patterns in the Africa rice (O. glaberrima and O. barthii): the more typical ‘Africa pattern’ characterized by the novel PSTOL1 allele and partial presence of the Pup1-specific INDEL region, but general absence of 90 kb of a region upstream of PSTOL1 (pattern G, Fig. 1); and the less common pattern (K) with Kasalath alleles across most of Pup1. Within O. sativa, the Kasalath (K) and Nipponbare (N) patterns could be distinguished as described earlier, but in addition a mixed pattern (m) with partial presence of Kasalath, O. glaberrima, and novel alleles was detected. These three patterns were already present in O. rufipogon and O. nivara, the wild ancestors of O. sativa. Results suggested that Pup1 locus variation was already a common feature within wild ancestors of cultivated Asian and African rice. Thus, divergence at Pup1 appears to predate domestication of rice.
Since the function of other genes within the Pup1 locus remains unclear, it would be desirable to transfer the entire Pup1 region from Kasalath into recipient varieties during marker-assisted selection. Thus, we propose using two foreground markers (K46-K and K20-K) in breeding programs aimed at introgressing Pup1.
Figure, table
-
Fig. 1. Characterization of the Pup1 locus in Nipponbare and Kasalath, and cultivated and wild rice genotypes. A main difference is the absence of a 90 kb INDEL region in Nipponbare containing OsPSTOL1 and 20 other Kasalath-specific genes.
-
Fig. 2. Amplification of OsPSTOL1 alleles using allele-specific markers for Kasalath and CG14 in single PCR (A), and duplex PCR system to detect both alleles in one reaction (B). (1: CG14, 2: IRAT, 3: NERICA16, 4: WAB56-50, 5: NERICA1, 6: NERICA10, 7: WAB181-18, 8: IDSA, 9: IR12979, 10: WAB56-104, 11: IAC165, 12: Nipponbare, 13: Kasalath)
- Affiliation
-
Japan International Research Center for Agricultural Sciences Crop, Livestock and Environment Division
- Classification
-
Administration A
- Research project
- Program name
- Term of research
-
FY 2014 (FY 2011-FY 2015)
- Responsible researcher
-
Wissuwa Matthias ( Crop, Livestock and Environment Division )
KAKEN Researcher No.: 90442722Tanaka Pariasca-Juan ( Crop, Livestock and Environment Division )
Chin Joong Hyoun ( International Rice Research Institute )
Dramé Khady Nani ( AfricaRice )
- ほか
- Publication, etc.
-
https://doi.org/10.1007/s00122-014-2306-y
J. Pariasca-Tanaka et al. (2014) Theor Appl Genet 127: 1387-1398.
- Japanese PDF
-
2014_B06_A3_ja.pdf553.23 KB
2014_B06_A4_ja.pdf495.13 KB
- English PDF
-
2014_B06_A3_en.pdf254.43 KB
2014_B06_A4_en.pdf247.77 KB
- Poster PDF
-
2014_B06_poster.pdf338.49 KB