Gene discovery of SPIKE -A unique gene from a rice landrace increases grain yield of Indica-type cultivars-
Description
Increasing crop production is essential for securing the future food supply in developing countries. Total spikelet number per panicle (TSN) is one of the most important traits to increase grain productivity in rice (Oryza sativa L.). We previously reported the detection of qTSN4, a QTL for increasing TSN on the long arm of chromosome 4 derived from new plant type (NPT) cultivars with the genetic background of IR64 (Refer the Research Highlight in 2012). In this study, we attempted to clone the gene for qTSN4. We further conducted characterization of agronomic traits of a near isogenic line (NIL) with the gene. To understand the effect of the genein different genetic backgrounds, we introduced it into six indica cultivars popular in South and Southeast Asian countries by marker assisted selection.
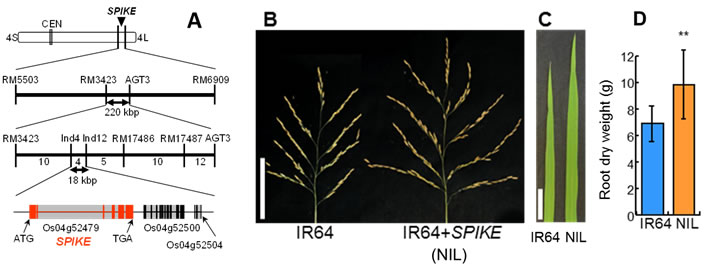
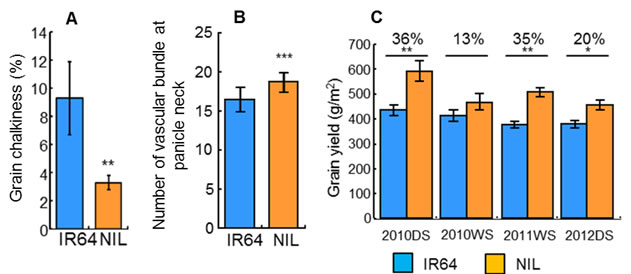
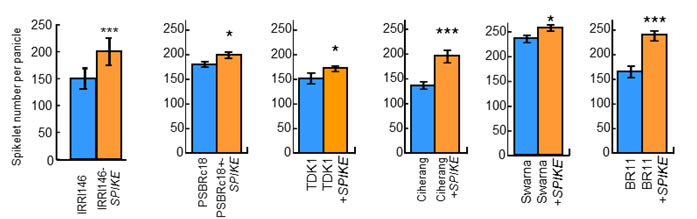
We successfully identified a causative gene for qTSN4, designated here as SPIKE (SPIKELET NUMBER) by map based cloning using 7996 BC4F3 plants of an NPT cultivar as a donor and IR64 as a recurrent parent (Fig. 1A). NIL for SPIKE had higher TSN (Fig. 1B), wider flag leaf (Fig. 1C) and heavier root dry weight (Fig. 1D) than those of IR64. Rate of filled grain were also higher, but panicle number per plant and 1000-grain weight were lower in the NIL (data not shown). Notably, the grain appearance of the NIL was significantly improved (Fig. 2A), presumably owing to a strengthening of assimilate supply to the larger number of spikelets by an increase in vascular bundle number (Fig. 2B). Consequently, the grain yield of the NIL was consistently higher by 13-36% than that of IR64 over four cropping seasons, significantly so in three of the four seasons (Fig. 2C). SPIKE also increased TSN in six cultivars popular in South and Southeast Asia (Fig. 3), confirming its effectiveness in various genetic backgrounds.
SPIKE would be a unique gene to increase grain yields of indica cultivars in South and Southeast Asia through molecular marker-assisted breeding, thus contributing to food security in these regions.
Figure, table
-
Fig. 1 Chromosomal location of SPIKE (A), panicle architecture (B), flag leaf width (C) and root dry weight (D). Values are mean ± SD. **Significant at 1% level by t-test.
-
Fig. 2 Percentages of grain chalkiness (A), number of vascular bundle at panicle neck (B) and grain yield (C). Values are mean ± SD (A, B) or SE (C). Significant at ***0.1%, **1% and *5% level by t-test.
-
Fig. 3 Total spikelet number per panicle between Indica-type cultivars with and without SPIKE. Values are mean ± SE. Significant at ***0.1%, **1% and *5% level by t-test.
- Affiliation
-
Japan International Research Center for Agricultural Sciences Biological Resources and Post-harvest Division
- Classification
-
Administration A
- Research project
- Program name
- Term of research
-
FY 2013 (FY 2005-FY 2013)
- Responsible researcher
-
Kobayashi Nobuya ( Institute of Crop Science, NARO )
KAKEN Researcher No.: 70252799Fujita Daisuke ( Institute of Crop Science, NARO )
KAKEN Researcher No.: 80721274Tagle Analiza G. ( International Rice Research Institute )
Koide Yohei ( Research Fellow of the Japan Society for the Promotion of Science )
KAKEN Researcher No.: 70712008Sasaki Kazuhiro ( University of Tokyo )
KAKEN Researcher No.: 70513688Gannaban Ritchel B. ( International Rice Research Institute )
Fukuta Yoshimichi ( Tropical Agriculture Research Front )
Ishimaru Tsutomu ( Biological Resources and Post-harvest Division )
- ほか
- Publication, etc.
-
https://doi.org/10.1073/pnas.1310790110
Fujita, D. et al. (2013) PNAS, 110: 20431-20436
- Japanese PDF
-
2013_A03_A3_ja.pdf294.43 KB
2013_A03_A4_ja.pdf423.12 KB
- English PDF
-
2013_A03_A3_en.pdf282.44 KB
2013_A03_A4_en.pdf211.52 KB
- Poster PDF
-
2013_A03_poster.pdf296.54 KB