A new international system of differentiating races of blast disease by using LTH monogenic lines in rice
Description
A new systematic, expandable method that allows easy understanding of the relationships between blast races and resistance genes is proposed for building up an international standard designation and classification system of blast races. Blast races were characterized by their reactions to 26 LTH (Lijiangxintuanheigu) monogenic lines used for targeting 23 resistance genes, which were divided into five groups, (1) LTH, IRBLa-A, IRBLsh-S, IRBLb-B, and IRBLt-K59, (2) three lines of Pii locus region, (3) seven lines of Pik region, (4) four lines of Piz region, and (5) seven lines of Pita region. Each group consists of one to three varieties, which were allocated with three differential lines (genes) in each and applied with codes 1, 2, and 4, for compatible reactions with blast isolates, respectively. Each blast race is characterized by the sum of codes in combinations of three varieties’ reactions in each unit using the Gilmour method for plant pathogen nomenclature. In the case of all compatible reactions to differential lines, the race No. 73-i7-k177-z17-ta733 was designated.
Evaluation and selection of blast isolate to monogenic lines
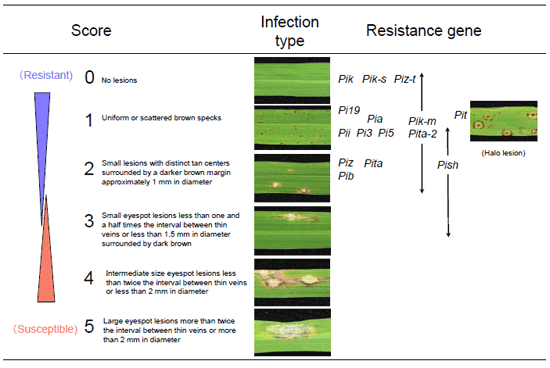
The 26 monogenic representative lines for 23 resistance genes, Pia, Pib, Pii, Pik, Pik-h, Pik-m, Pik-p, Pik-s, Pish, Pit, Pita, Pita-2, Piz, Piz-t, Piz-5, Pi1, Pi3, Pi5(t), Pi7(t), Pi9(t), Pi12(t), Pi19(t), and Pi20(t), including LTH as a susceptible check variety, were used. The responses of each LTH monogenic line to blast fungal isolates were recorded according to the infection type (generally, 0–2 resistant; 3–5 susceptible) in accordance with the IRRI Standard Evaluation System (SES, 1996) (Fig. 1).
Grouping of differential varieties
Twenty of the LTH monogenic lines are multiallelic genes in four genetic loci. The LTH monogenic lines, having genes presumed to be multiallelic or closely linked in the Pii, Pik, Piz, and Pita locus, were divided into groups, respectively, and arranged in accordance with the Gilmour method. Each race code number has the following five parts divided by a hyphen (i.e., 1st-2nd-3rd-4th-5th). The first part of the race code number has a two-digit number and is composed of LTH, and IRBLa-A in the ones place, and IRBLsh-S, IRBLb-B and IRBLt-K59 in the tens place. The second part has a one-digit number and is composed of IRBLi-F5, IRBL3-CP4, and IRBL5-M, which are multiallelic or closely linked. The third part has a three-digit number and is composed of IRBLk-Ka, IRBLkp-K60 and IRBL7-M in the ones place, IRBLkm-Ts, IRBL1-CL and IRBLkh-K3 in the tens place, and IRBLks-S, in the hundreds place, which are multiallelic or closely linked. The fourth part has a two-digit number and is composed IRBLz-Fu, IRBLz5-CA, and IRBLzt-T in the ones place, and IRBL9-W in the tens place, which are multiallelic or closely linked. The fifth part has a three-digit number and is composed IRBL19-A, IRBL20-IR24 in the ones place, IRBLta-K1, IRBLta-CP1 in the tens place, and IRBLta2-Pi, IRBLta2-Re, IRBL12-M in the hundreds place, which are multiallelic or closely linked.
Classifications
To clearly delineate the relationship between race code number and resistance gene, the system of nomenclature of blast fungus races employed the Gilmour’s method. This nomenclature method is excellent in terms of regularity, flexibility and suitability for a differential system composed of many varieties. The selected LTH monogenic lines were divided into groups with three lines each, and each of the three lines was given the code numbers 1, 2, and 4, respectively. The total of the code numbers of the three LTH monogenic lines, which are compatible to blast fungus, show the race number.
Designation system
A race number is shown in the total of the code numbers of monogenic lines, which are compatible to a specific blast fungus. To indicate which of each part is the Pii, Pik, Piz and Pita locus, the symbols i, k, z and ta, which show the locus name, were marked in front of each multiallelic part, respectively. For example, when a blast isolate is pathogenic to all the differential varieties, the race number of the isolate is shown as “73-i7-k177-z17-ta773”. This system offers sufficient range for regularity, flexibility and expansion to take in new resistance varieties to be identified in the future. Usually, when the number of differential varieties are increased along with the introduction of new resistance genes, extensibility is poor.
This study was conducted under the research projects “Blast Research Network for Stable Rice Production” funded by JIRCAS, and the IRRI-Japan Collaborative Research Project Phases IV and V, commissioned to IRRI by the Ministry of Agriculture, Forestry and Fisheries (MAFF) of Japan.
Figure, table
-
Fig. 1. Criteria for evaluating infection types of blast races.
- Affiliation
-
Japan International Research Center for Agricultural Sciences Biological Resources Division
-
National Institute of Agrobiological Sciences
- Classification
-
Technical A
- Term of research
-
FY2007(FY2006~2010)
- Responsible researcher
-
HAYASHI Nagao ( National Institute of Agrobiological Sciences )
FUKUTA Yoshimichi ( Biological Resources Division )
- ほか
- Publication, etc.
-
Hayashi, N. (2006) System for designation of blast races using international rice blast line -Proposal of a new method-. Workshop on a differential system for blast resistance for a stable rice production environment. p6-7, IRRI, Los Banos.
Hayashi and Fukuta (2007) Abstracts of Phytopathological Society of Japan Annual Meeting, p74, Lecture No. 223.
Hayashi et al. (2008) Abstracts of Phytopathological Society of Japan Annual Meeting, p71, Lecture No. 213 4. 26-28
Hayashi, N., Fukuta, Y. (2008) New designation system for pathogenic race of rice blast fungus using LTH monogenic lines in rice. JIRCAS Workshop In Identification and characterization of blast race and resistance gene based on differential system, 4-5.
Hayashi, N., Kobayashi, N., Vera Cruz, C.M., Fukuta, Y. (2009) Protocols for the sampling of diseased specimens and evaluation of blast disease in rice. JIRCAS Working Report 63,17-33.
Development and Characterization of Blast Resistance Using Differential Varieties in Rice
- Japanese PDF
-
2008_seikajouhou_A4_ja_Part8.pdf515.31 KB